MultiLR: Prediction + inferential models
STA 210 - Summer 2022
Yunran Chen
Welcome
Topics
- Predictions
- Model selection
- Checking conditions
Computational setup
NHANES Data
- National Health and Nutrition Examination Survey is conducted by the National Center for Health Statistics (NCHS).
- The goal is to “assess the health and nutritional status of adults and children in the United States”.
- This survey includes an interview and a physical examination.
Variables
Goal: Use a person’s age and whether they do regular physical activity to predict their self-reported health rating.
Outcome:
HealthGen: Self-reported rating of participant’s health in general. Excellent, Vgood, Good, Fair, or Poor.Predictors:
Age: Age at time of screening (in years). Participants 80 or older were recorded as 80.PhysActive: Participant does moderate to vigorous-intensity sports, fitness or recreational activities.
The data
Rows: 6,465
Columns: 5
$ HealthGen <fct> Good, Good, Good, Good, Vgood, Vgood, Vgood, Vgood, Vgood, …
$ Age <int> 34, 34, 34, 49, 45, 45, 45, 66, 58, 54, 50, 33, 60, 56, 56,…
$ PhysActive <fct> No, No, No, No, Yes, Yes, Yes, Yes, Yes, Yes, Yes, No, No, …
$ Education <fct> High School, High School, High School, Some College, Colleg…
$ obs_num <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, …Model in R
Model summary
# A tibble: 12 × 6
y.level term estimate std.error statistic p.value
<chr> <chr> <dbl> <dbl> <dbl> <dbl>
1 Vgood (Intercept) 1.27 0.154 8.23 1.80e-16
2 Vgood Age -0.0000361 0.00259 -0.0139 9.89e- 1
3 Vgood PhysActiveYes -0.332 0.0949 -3.50 4.72e- 4
4 Good (Intercept) 1.99 0.150 13.3 2.81e-40
5 Good Age -0.00304 0.00256 -1.19 2.35e- 1
6 Good PhysActiveYes -1.01 0.0921 -11.0 4.80e-28
7 Fair (Intercept) 1.03 0.174 5.94 2.89e- 9
8 Fair Age 0.00113 0.00302 0.373 7.09e- 1
9 Fair PhysActiveYes -1.66 0.109 -15.2 4.14e-52
10 Poor (Intercept) -1.34 0.299 -4.47 7.65e- 6
11 Poor Age 0.0193 0.00505 3.83 1.30e- 4
12 Poor PhysActiveYes -2.67 0.236 -11.3 1.20e-29Predictions
Calculating probabilities
For categories \(2,\ldots,K\), the probability that the \(i^{th}\) observation is in the \(j^{th}\) category is
\[ \hat{\pi}_{ij} = \frac{e^{\hat{\beta}_{0j} + \hat{\beta}_{1j}x_{i1} + \dots + \hat{\beta}_{pj}x_{ip}}}{1 + \sum\limits_{k=2}^K e^{\hat{\beta}_{0k} + \hat{\beta}_{1k}x_{i1} + \dots \hat{\beta}_{pk}x_{ip}}} \]
For the baseline category, \(k=1\), we calculate the probability \(\hat{\pi}_{i1}\) as
\[ \hat{\pi}_{i1} = 1- \sum\limits_{k=2}^K \hat{\pi}_{ik} \]
Predicted health rating
We can use our model to predict a person’s perceived health rating given their age and whether they exercise.
# A tibble: 6,465 × 11
HealthGen Age PhysActive Education obs_num .pred_class .pred_Excellent
<fct> <int> <fct> <fct> <int> <fct> <dbl>
1 Good 34 No High School 1 Good 0.0687
2 Good 34 No High School 2 Good 0.0687
3 Good 34 No High School 3 Good 0.0687
4 Good 49 No Some College 4 Good 0.0691
5 Vgood 45 Yes College Grad 5 Vgood 0.155
6 Vgood 45 Yes College Grad 6 Vgood 0.155
7 Vgood 45 Yes College Grad 7 Vgood 0.155
8 Vgood 66 Yes Some College 8 Vgood 0.157
9 Vgood 58 Yes College Grad 9 Vgood 0.156
10 Fair 54 Yes 9 - 11th Grade 10 Vgood 0.156
# … with 6,455 more rows, and 4 more variables: .pred_Vgood <dbl>,
# .pred_Good <dbl>, .pred_Fair <dbl>, .pred_Poor <dbl>Actual vs. predicted health rating
For each observation, the predicted perceived health rating is the category with the highest predicted probability.
# A tibble: 6,465 × 6
.pred_class .pred_Excellent .pred_Vgood .pred_Good .pred_Fair .pred_Poor
<fct> <dbl> <dbl> <dbl> <dbl> <dbl>
1 Good 0.0687 0.243 0.453 0.201 0.0348
2 Good 0.0687 0.243 0.453 0.201 0.0348
3 Good 0.0687 0.243 0.453 0.201 0.0348
4 Good 0.0691 0.244 0.435 0.205 0.0467
5 Vgood 0.155 0.393 0.359 0.0868 0.00671
6 Vgood 0.155 0.393 0.359 0.0868 0.00671
7 Vgood 0.155 0.393 0.359 0.0868 0.00671
8 Vgood 0.157 0.400 0.342 0.0904 0.0102
9 Vgood 0.156 0.397 0.349 0.0890 0.00872
10 Vgood 0.156 0.396 0.352 0.0883 0.00804
# … with 6,455 more rowsConfusion matrix
health_conf <- health_aug %>%
count(HealthGen, .pred_class, .drop = FALSE) %>%
pivot_wider(names_from = .pred_class, values_from = n)
health_conf# A tibble: 5 × 6
HealthGen Excellent Vgood Good Fair Poor
<fct> <int> <int> <int> <int> <int>
1 Excellent 0 528 210 0 0
2 Vgood 0 1341 743 0 0
3 Good 0 1226 1316 0 0
4 Fair 0 296 625 0 0
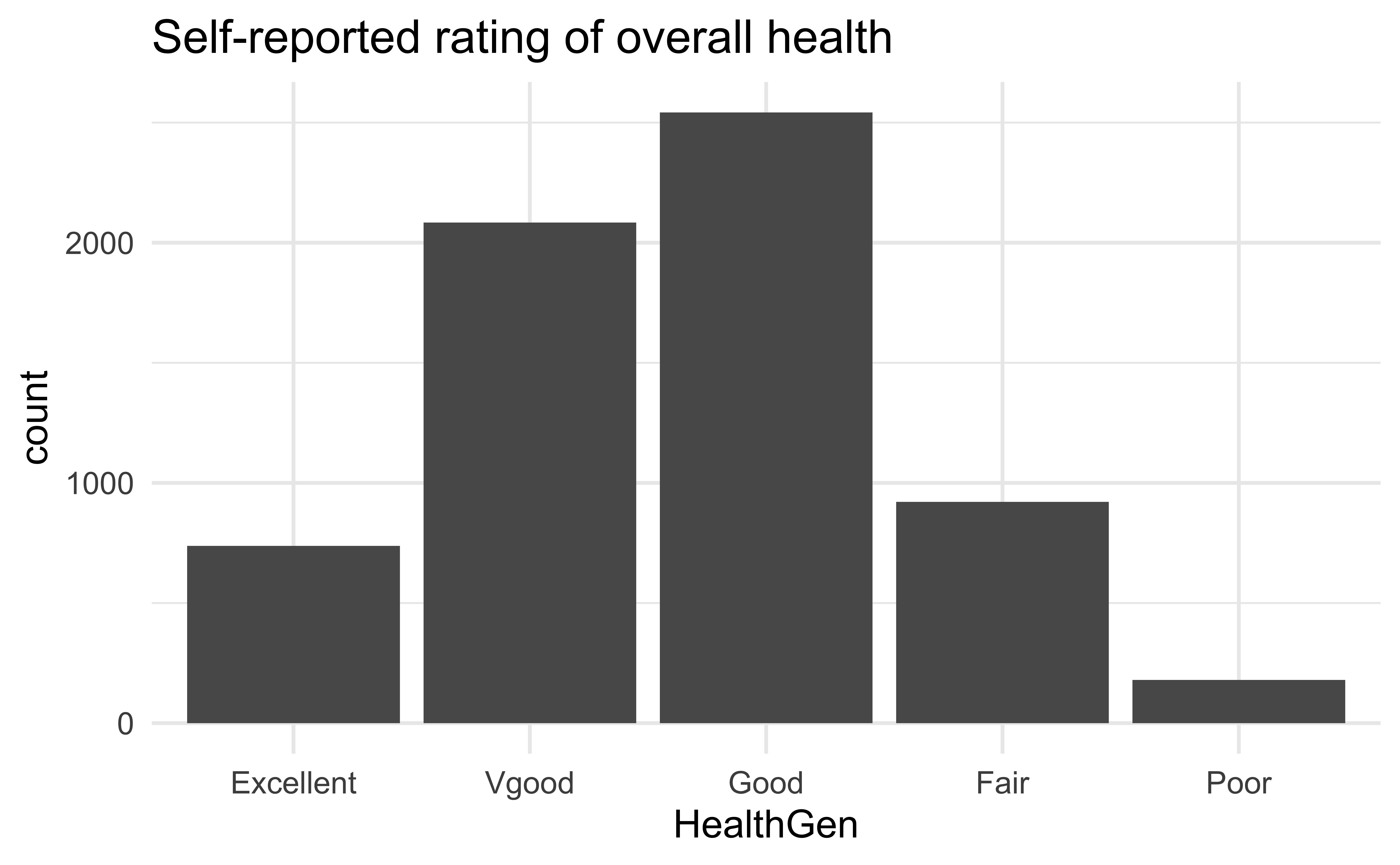
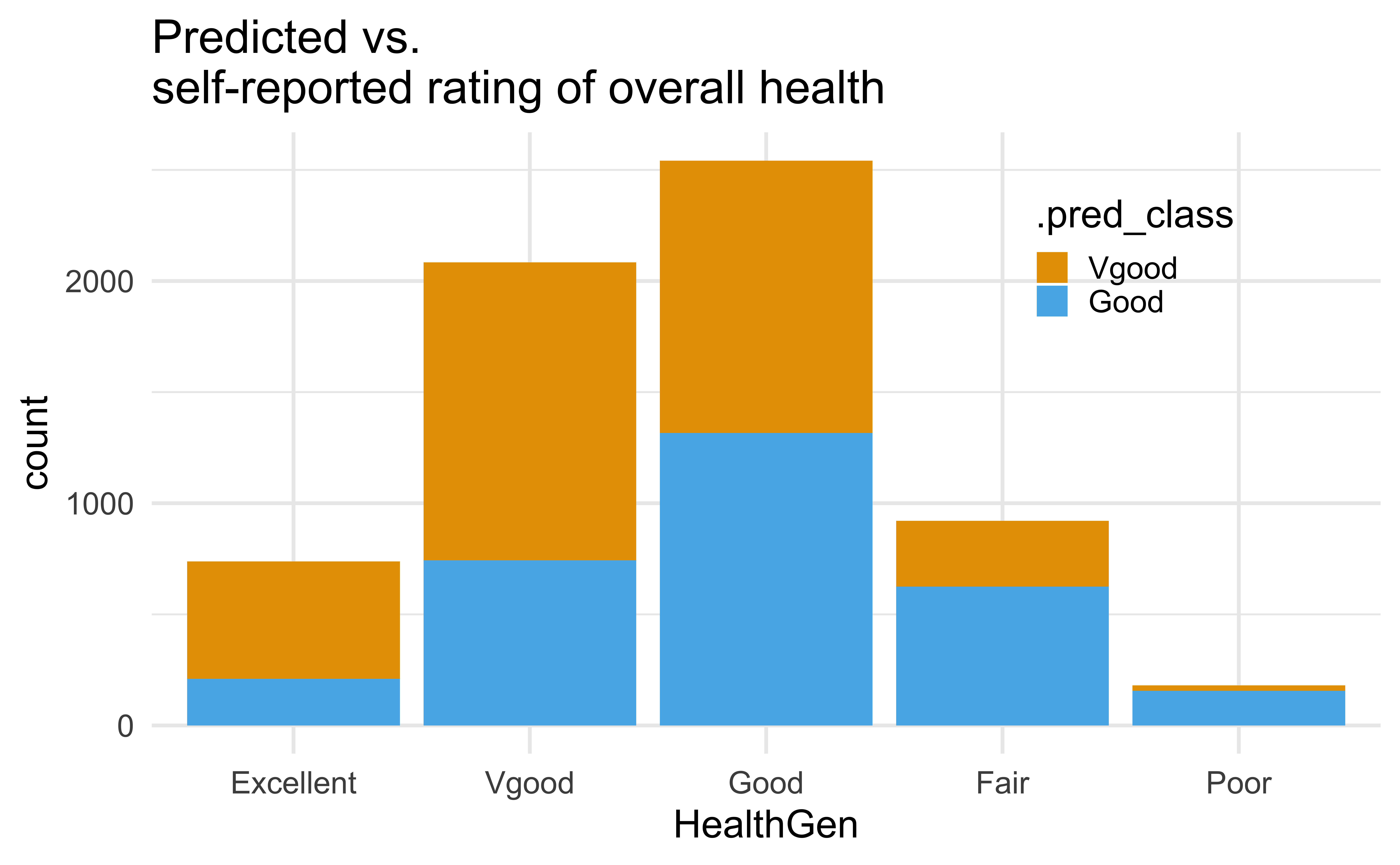
5 Poor 0 24 156 0 0Actual vs. predicted health rating
Why do you think no observations were predicted to have a rating of “Excellent”, “Fair”, or “Poor”?
Will discuss this later in the model diagnosis.
Model selection for inference
Comparing nested models
- Suppose there are two models:
- Reduced model includes predictors \(x_1, \ldots, x_q\)
- Full model includes predictors \(x_1, \ldots, x_q, x_{q+1}, \ldots, x_p\)
- We want to test the following hypotheses:
- \(H_0: \beta_{q+1} = \dots = \beta_p = 0\)
- \(H_A: \text{ at least 1 }\beta_j \text{ is not } 0\)
- To do so, we will use the drop-in-deviance test (very similar to logistic regression)
Add Education to the model?
- We consider adding the participants’
Educationlevel to the model.- Education takes values
8thGrade,9-11thGrade,HighSchool,SomeCollege, andCollegeGrad
- Education takes values
- Models we’re testing:
- Reduced model:
Age,PhysActive - Full model:
Age,PhysActive,Education
- Reduced model:
\[ \begin{align} &H_0: \beta_{9-11thGrade} = \beta_{HighSchool} = \beta_{SomeCollege} = \beta_{CollegeGrad} = 0\\ &H_a: \text{ at least one }\beta_j \text{ is not equal to }0 \end{align} \]
Add Education to the model?
reduced_fit <- multinom_reg() %>%
set_engine("nnet") %>%
fit(HealthGen ~ Age + PhysActive,
data = nhanes_adult)
full_fit <- multinom_reg() %>%
set_engine("nnet") %>%
fit(HealthGen ~ Age + PhysActive + Education,
data = nhanes_adult)
reduced_fit <- repair_call(reduced_fit, data = nhanes_adult)
full_fit <- repair_call(full_fit, data = nhanes_adult)Add Education to the model?
| Model | Resid. df | Resid. Dev | Test | Df | LR stat. | Pr(Chi) |
|---|---|---|---|---|---|---|
| Age + PhysActive | 25848 | 16994.23 | NA | NA | NA | |
| Age + PhysActive + Education | 25832 | 16505.10 | 1 vs 2 | 16 | 489.132 | 0 |
At least one coefficient associated with Education is non-zero. Therefore, we will include Education in the model.
If you want to apply anova to the fitted model from workflow, you have to apply fitted_obj %>% extract_fit_engine() to obtain fitted object that can be put into anova().
Model with Education
# A tibble: 28 × 8
y.level term estimate std.error statistic p.value conf.low conf.high
<chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 Vgood (Intercept) 5.82e-1 0.301 1.93 5.36e- 2 -0.00914 1.17
2 Vgood Age 1.12e-3 0.00266 0.419 6.75e- 1 -0.00411 0.00634
3 Vgood PhysActiveY… -2.64e-1 0.0985 -2.68 7.33e- 3 -0.457 -0.0711
4 Vgood Education9 … 7.68e-1 0.308 2.49 1.27e- 2 0.164 1.37
5 Vgood EducationHi… 7.01e-1 0.280 2.51 1.21e- 2 0.153 1.25
6 Vgood EducationSo… 7.88e-1 0.271 2.90 3.71e- 3 0.256 1.32
7 Vgood EducationCo… 4.08e-1 0.268 1.52 1.28e- 1 -0.117 0.933
8 Good (Intercept) 2.04e+0 0.272 7.51 5.77e-14 1.51 2.57
9 Good Age -1.72e-3 0.00263 -0.651 5.15e- 1 -0.00688 0.00345
10 Good PhysActiveY… -7.58e-1 0.0961 -7.88 3.16e-15 -0.946 -0.569
11 Good Education9 … 3.60e-1 0.275 1.31 1.90e- 1 -0.179 0.899
12 Good EducationHi… 8.52e-2 0.247 0.345 7.30e- 1 -0.399 0.569
13 Good EducationSo… -1.13e-2 0.239 -0.0472 9.62e- 1 -0.480 0.457
14 Good EducationCo… -8.91e-1 0.236 -3.77 1.65e- 4 -1.35 -0.427
15 Fair (Intercept) 2.12e+0 0.288 7.35 1.91e-13 1.55 2.68
16 Fair Age 3.35e-4 0.00312 0.107 9.14e- 1 -0.00578 0.00645
17 Fair PhysActiveY… -1.19e+0 0.115 -10.4 3.50e-25 -1.42 -0.966
18 Fair Education9 … -2.24e-1 0.279 -0.802 4.22e- 1 -0.771 0.323
19 Fair EducationHi… -8.32e-1 0.252 -3.31 9.44e- 4 -1.33 -0.339
20 Fair EducationSo… -1.34e+0 0.246 -5.46 4.71e- 8 -1.82 -0.861
21 Fair EducationCo… -2.51e+0 0.253 -9.91 3.67e-23 -3.00 -2.01
22 Poor (Intercept) -2.00e-1 0.411 -0.488 6.26e- 1 -1.01 0.605
23 Poor Age 1.79e-2 0.00509 3.53 4.21e- 4 0.00797 0.0279
24 Poor PhysActiveY… -2.27e+0 0.242 -9.38 6.81e-21 -2.74 -1.79
25 Poor Education9 … -3.60e-1 0.353 -1.02 3.08e- 1 -1.05 0.332
26 Poor EducationHi… -1.15e+0 0.334 -3.44 5.86e- 4 -1.81 -0.494
27 Poor EducationSo… -1.07e+0 0.316 -3.40 6.77e- 4 -1.69 -0.454
28 Poor EducationCo… -2.32e+0 0.366 -6.34 2.27e-10 -3.04 -1.60 Compare NHANES models using AIC
Reduced model:
We can also use BIC to compare model. Since the glance() does not provide us the BIC. For this class, we just use AIC.
Checking conditions for inference
Conditions for inference
We want to check the following conditions for inference for the multinomial logistic regression model:
Linearity: Is there a linear relationship between the log-odds and the predictor variables?
Randomness: Was the sample randomly selected? Or can we reasonably treat it as random?
Independence: Are the observations independent?
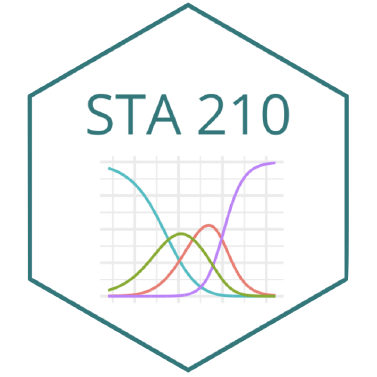
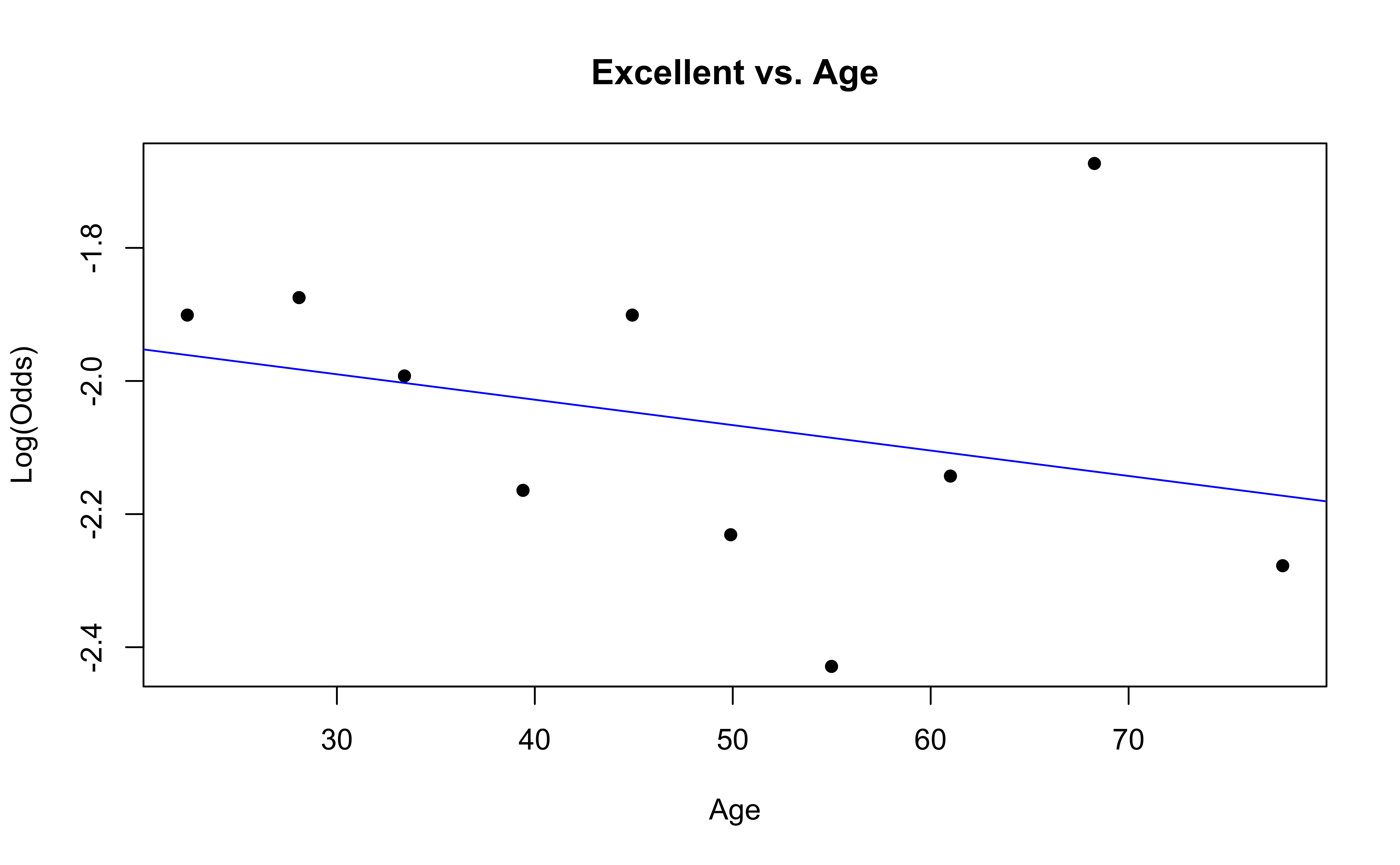
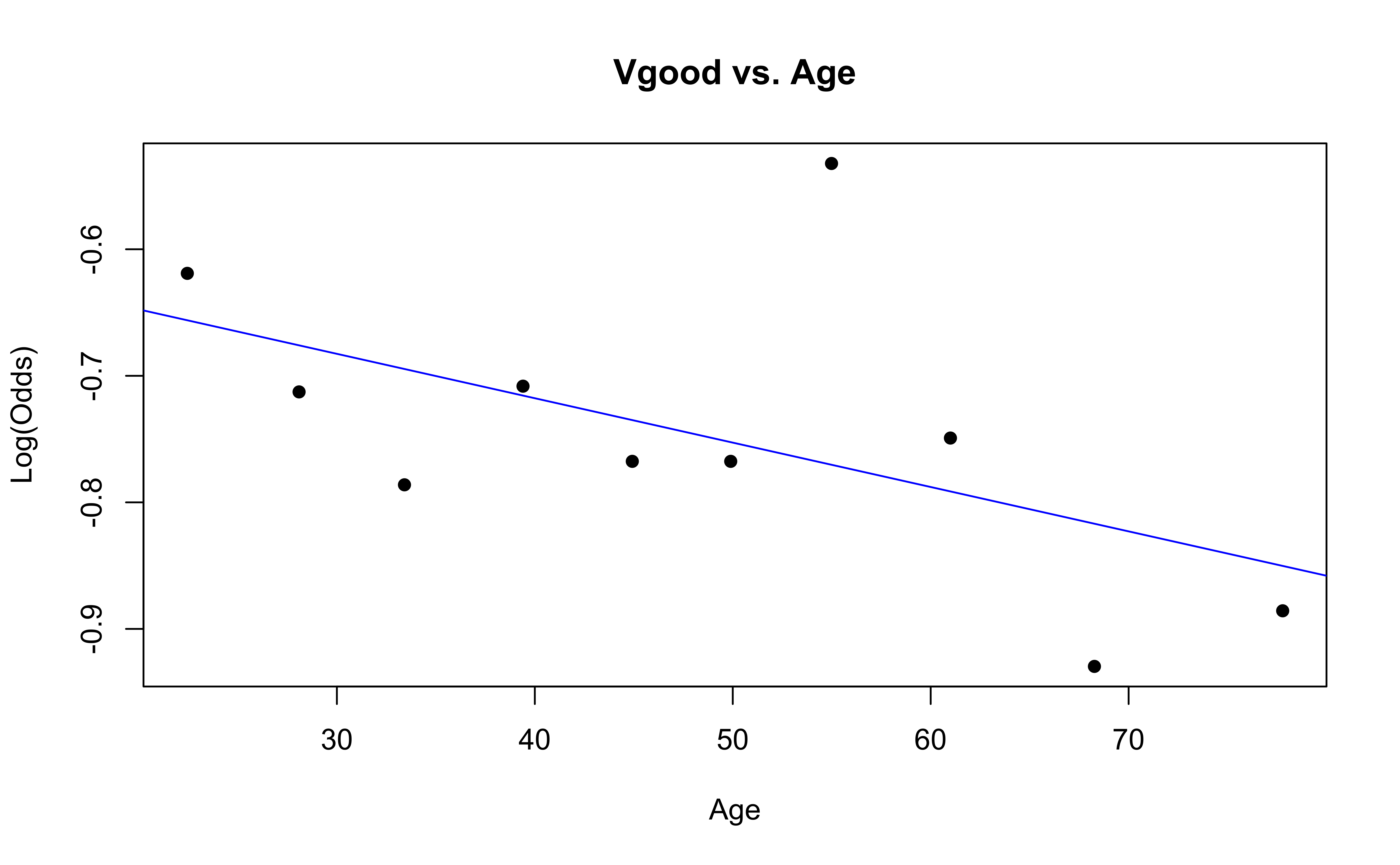
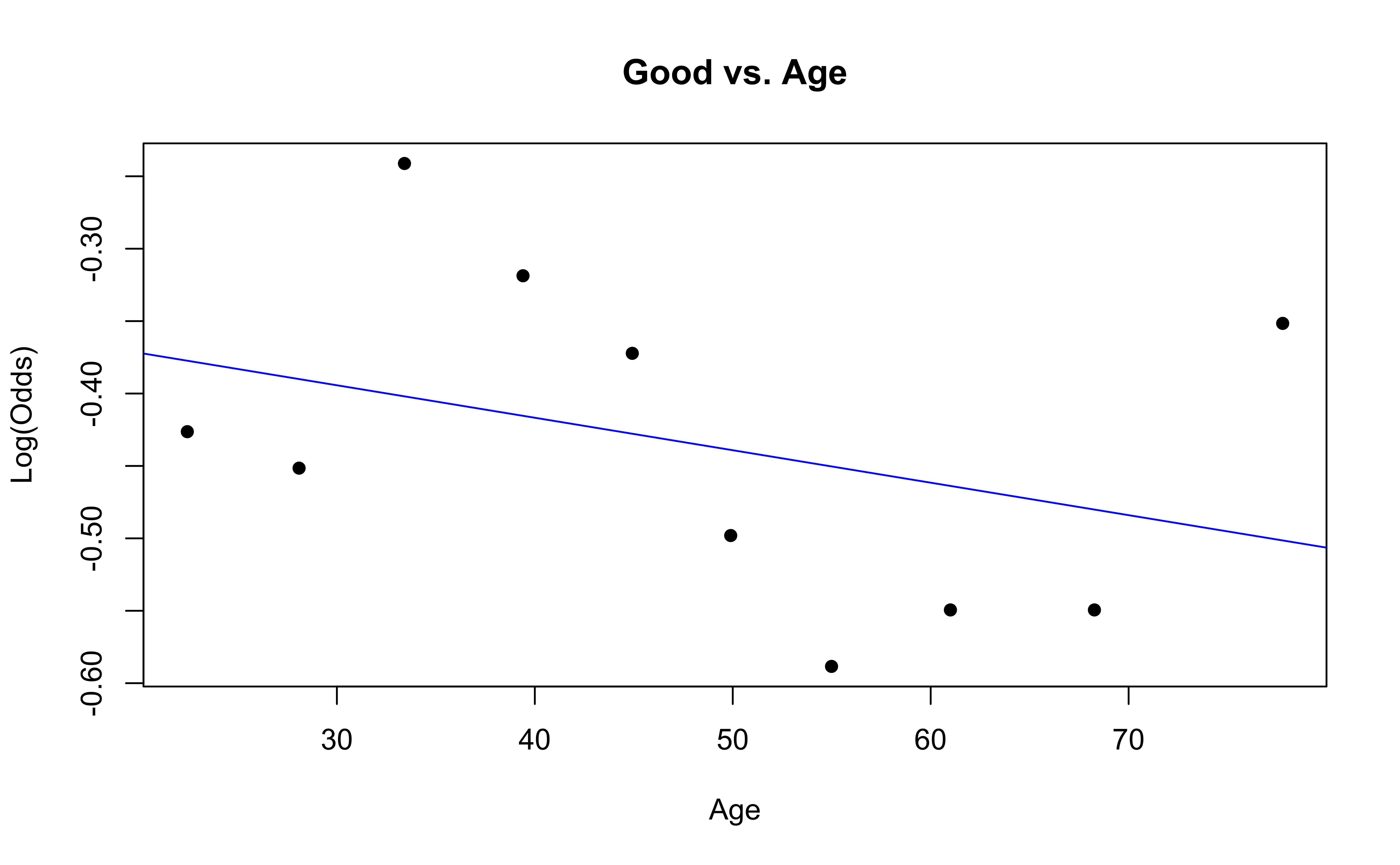
Checking linearity
Similar to logistic regression, we will check linearity by examining empirical logit plots between each level of the response and the quantitative predictor variables.
nhanes_adult <- nhanes_adult %>%
mutate(
Excellent = factor(if_else(HealthGen == "Excellent", "1", "0")),
Vgood = factor(if_else(HealthGen == "Vgood", "1", "0")),
Good = factor(if_else(HealthGen == "Good", "1", "0")),
Fair = factor(if_else(HealthGen == "Fair", "1", "0")),
Poor = factor(if_else(HealthGen == "Poor", "1", "0"))
)Checking linearity
emplogitplot1: numerical predictor
emplogitplot2: numerical + categorical
Checking linearity
Checking linearity
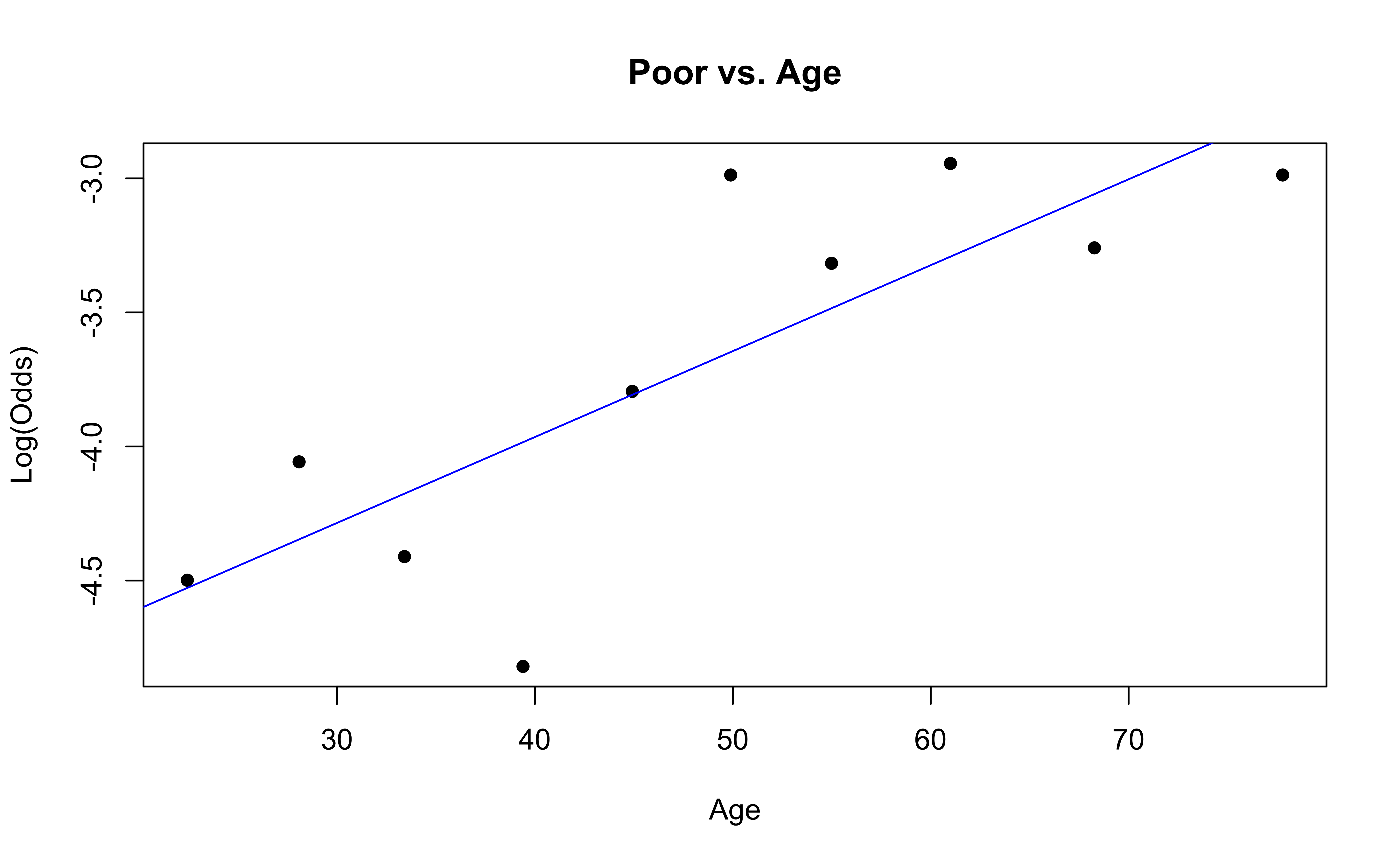
The linearity condition is not satisfied. If we recall the previous confusion matrix, the fitted model only provide two fitted class: very good and good, which may due to the unreasonable model assumption.
Checking randomness
We can check the randomness condition based on the context of the data and how the observations were collected.
Was the sample randomly selected?
If the sample was not randomly selected, ask whether there is reason to believe the observations in the sample differ systematically from the population of interest.
✅ The randomness condition is satisfied. We do not have reason to believe that the participants in this study differ systematically from adults in the U.S..
Checking independence
We can check the independence condition based on the context of the data and how the observations were collected.
Independence is most often violated if the data were collected over time or there is a strong spatial relationship between the observations.
✅ The independence condition is satisfied. It is reasonable to conclude that the participants’ health and behavior characteristics are independent of one another.
Recap
- Predictions
- Model selection for inference
- Checking conditions for inference